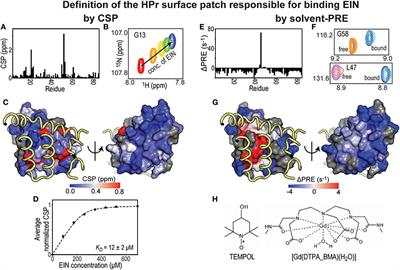
Quantitative Interpretation of Solvent Paramagnetic Relaxation for Probing Protein–Cosolute Interactions | Journal of the American Chemical Society

Theory and Applications of Nitroxide-based Paramagnetic Cosolutes for Probing Intermolecular and Electrostatic Interactions on Protein Surfaces | Journal of the American Chemical Society

Atomic resolution protein allostery from the multi-state structure of a PDZ domain | Nature Communications

A large data set comparison of protein structures determined by crystallography and NMR: Statistical test for structural differences and the effect of crystal packing - Andrec - 2007 - Proteins: Structure, Function,

Bioinformatic Analysis and Biophysical Characterization Reveal Structural Disorder in G0S2 Protein | ACS Omega

Systematic Exploration of Protein Conformational Space Using a Distance Geometry Approach | Journal of Chemical Information and Modeling
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/4-Table3-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar

Frontiers | An Integrative Approach to Determine 3D Protein Structures Using Sparse Paramagnetic NMR Data and Physical Modeling
Determination of three-dimensional structures of proteins in solution by nuclear magnetic resonance spectroscopy

Functional protein dynamics on uncharted time scales detected by nanoparticle-assisted NMR spin relaxation | Science Advances

Sequence‐specific determination of protein and peptide concentrations by absorbance at 205 nm - Anthis - 2013 - Protein Science - Wiley Online Library

Table 1 from Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar

Quantitative analysis of sterol-modulated monomer–dimer equilibrium of the β1-adrenergic receptor by DEER spectroscopy | PNAS

Molecules | Free Full-Text | Patterns in Protein Flexibility: A Comparison of NMR “Ensembles”, MD Trajectories, and Crystallographic B-Factors

Quantitative Interpretation of Solvent Paramagnetic Relaxation for Probing Protein–Cosolute Interactions | Journal of the American Chemical Society

PDBcor: An automated correlation extraction calculator for multi-state protein structures - ScienceDirect

Chimeric single α-helical domains as rigid fusion protein connections for protein nanotechnology and structural biology - ScienceDirect